2015 – Identification of Proteins in Raw Materials and Finished Goods by LC/MS/MS
resources - Recent Conference Materials

UNPA Dietary Supplement Analytical Summit
HISTORY/CURRENT PRACTICE TO IDENTIFY PROTEINS
The total nitrogen content methods have long been used to assay proteins. The Kjehldahl and Dumas methods are most common. Interpretations of these results assume the source of the nitrogen comes from proteins. However, this assumption has been exploited in the past. In 2007, there was a scandal with melamine (it was nitrogen-rich, but not a protein) spiked into pet food, infant formula, etc. Today, Inexpensive amino acids can be added to increase nitrogen content. These methods do not distinguish between different types of proteins and are still accepted and widely used today. Other methods that are currently used to identify proteins, are Organoleptic (this is subjective), Relative comparison of amino acids by LC/UV (not selective for finished products), and Colorimetric/fluorescent kits (not selective; does not distinguish between types of proteins).
RAISING THE BAR FOR TESTING
Agencies are increasing activity to enforce exactly what is on labels, which is requiring more accurate and precise testing. Consumers are being influenced by recent events and becoming more knowledgeable. The current industry practice for analysis of protein testing is vulnerable to exposure in the future. And alternative approach to testing is to select a Specific Protein. The Sample Preparation would consist of breaking down the protein into smaller peptide fragments.
PROTEIN SELECTION, LIQUID CHROMATOGRAPHY
There are multiple proteins are present in samples. We selected a specific protein to monitor based on literature data. Our example used was SDS-Page of seed total protein in rice (glutelin chosen here). In sample preparation the Trypsin enzyme hydrolyzes proteins. Cleavage occurs mainly at the carboxyl side of the amino acids Lysine (K) and Arginine (R). Smaller tryptic peptide fragments result from cleavage. In liquid chromatography testing, peptides can be separated from each other and other components in the extract by liquid chromatography (Shimadzu Prominence). Peptides can be detected in a mass spectrometer (Sciex API4000 QTrap) based on their mass-to-charge (m/z) ratio in positive-ion mode. Peptides fragment in a collision cell and product ions are monitored. Typically, double-charge peptides provide the highest signal.
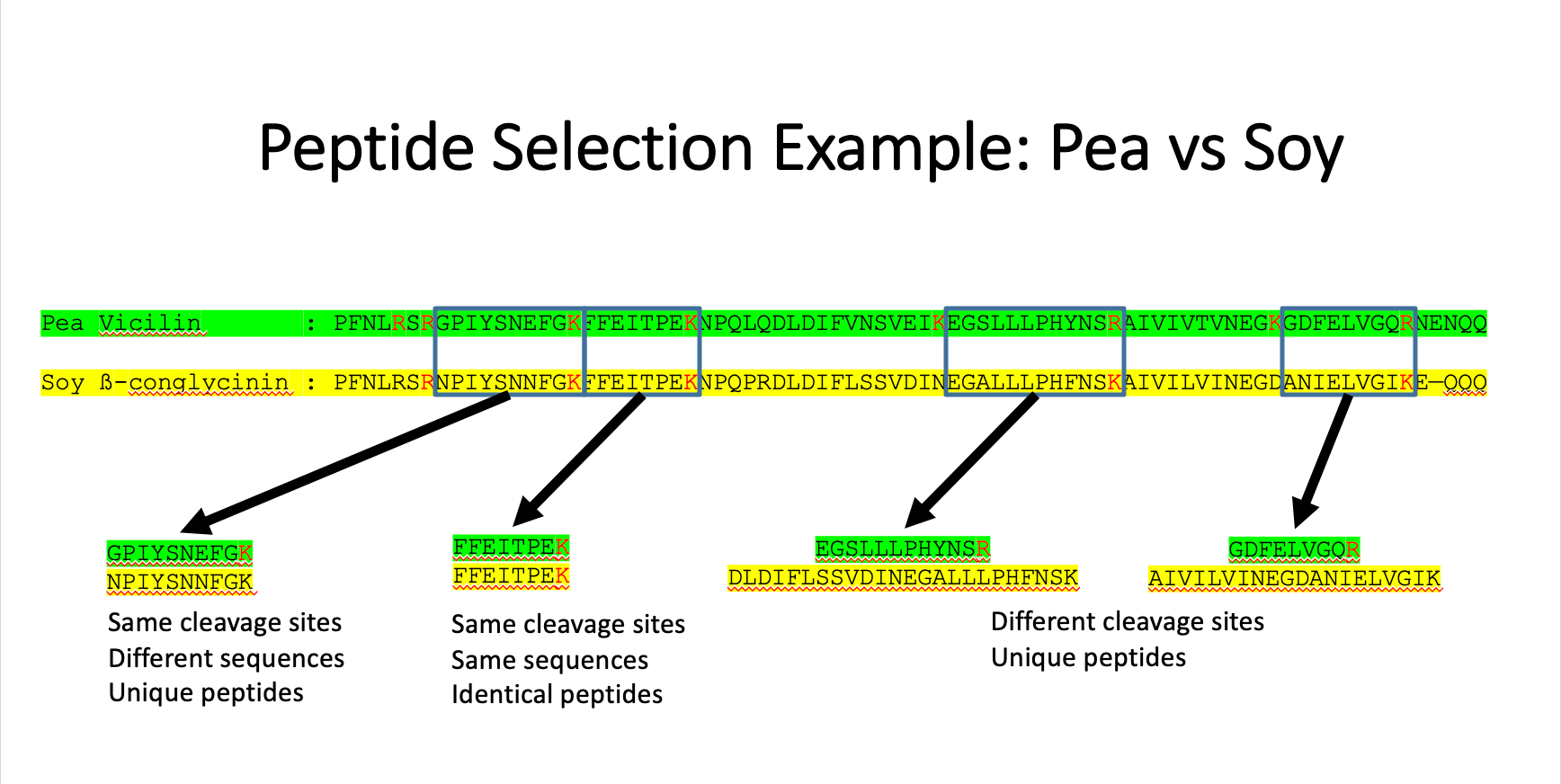
PEPTIDE SELECTION
Similar to DNA barcoding, only a specific region of the protein’s amino acid sequence is chosen. We selected unique tryptic peptides from the sequence. Multiple peptide MS transitions can be monitored simultaneously (three for each protein). See multiple examples below and in the presentation PDF.
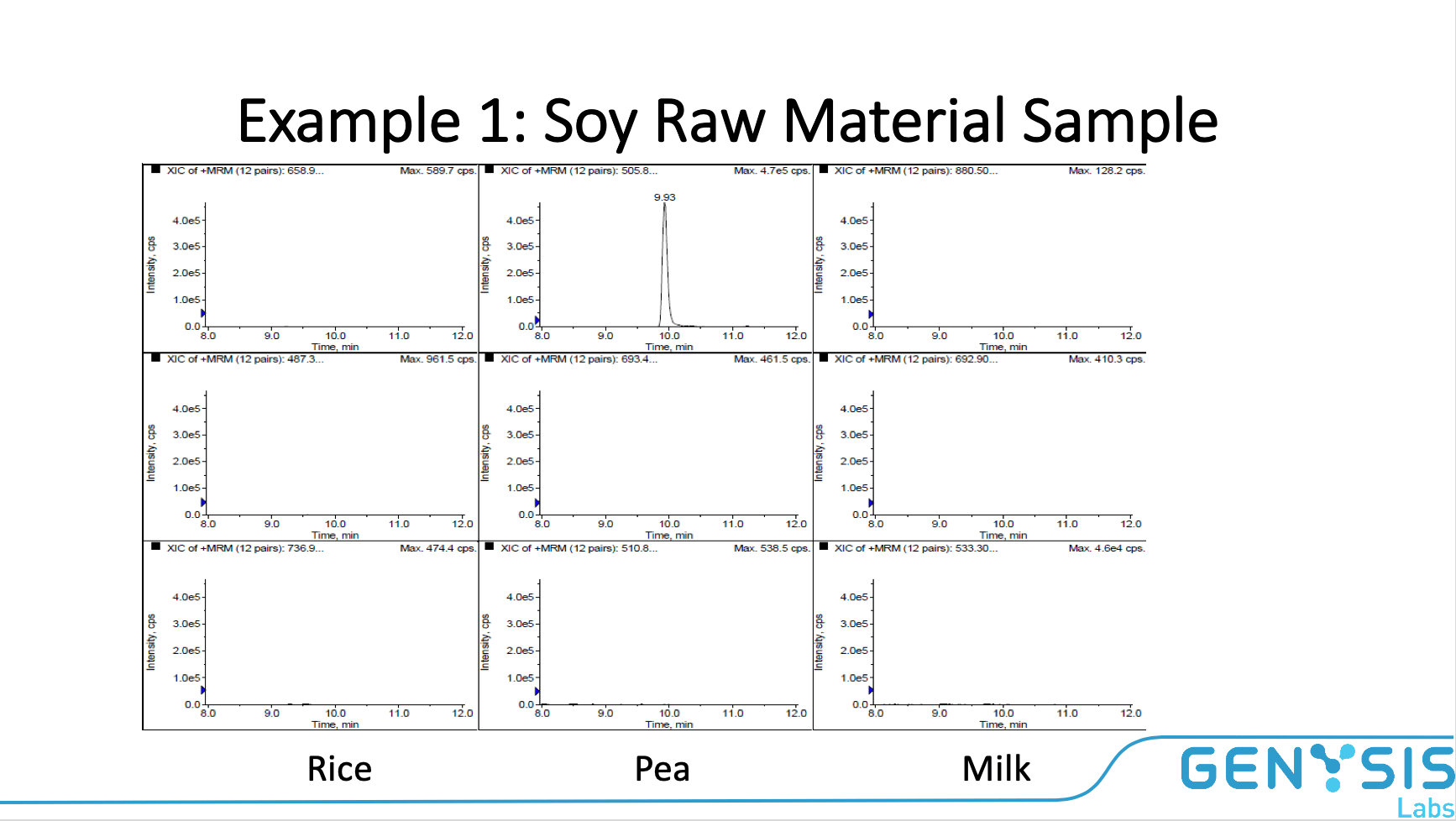
CONCLUSION
Proteins are generally stable through processing, which allows analysis on finished products as well as raw materials. Trypsin digest of samples followed by LC/MS/MS analysis provides conclusive protein identification. Assay is selective. We can monitor unique and specific peptides from specific proteins. The chromatographic retention time of peptides is the initial confirmation. Peaks using a mass transition of a fragmented peptide are monitored for additional confirmation. A peak in all three transitions is needed for confirmation. Positive (protein standards) and negative (blank samples) controls are additional confirmation.